4.3 Proteins
Learning Objectives
By the end of this section, you will be able to:
- Describe the functions proteins perform in the cell and in tissues
- Discuss the relationship between amino acids and proteins
- Explain the four levels of protein organization
- Describe the ways in which protein shape and function are linked
Proteins are one of the most abundant organic molecules in living systems and have the most diverse range of functions of all macromolecules. Proteins may be structural, regulatory, contractile, or protective. They may serve in transport, storage, or membranes; or they may be toxins or enzymes. Each cell in a living system may contain thousands of proteins, each with a unique function. Their structures, like their functions, vary greatly. They are all, however, amino acid polymers arranged in a linear sequence.
Types and Functions of Proteins
Proteins have different shapes and molecular weights. Table 4.1 lists the primary types and functions of proteins. Some proteins are globular in shape; whereas, others are fibrous in nature. For example, hemoglobin is a globular protein, but collagen, located in our skin, is a fibrous protein. Protein shape is critical to its function, and many different types of chemical bonds maintain this shape. Changes in temperature, pH, and exposure to chemicals may lead to permanent changes in the protein’s shape, leading to loss of function, or denaturation.
Enzymes, which living cells produce, are catalysts in biochemical reactions (like digestion) and are usually complex or conjugated proteins. Each enzyme is specific for the substrate (a reactant that binds to an enzyme) upon which it acts. The enzyme may help in breakdown, rearrangement, or synthesis reactions. We call enzymes that break down their substrates catabolic enzymes. Those that build more complex molecules from their substrates are anabolic enzymes, and enzymes that affect the rate of reaction are catalytic enzymes. Note that all enzymes increase the reaction rate and, therefore, are organic catalysts. An example of an enzyme is salivary amylase, which hydrolyzes its substrate amylose, a component of starch.
Hormones are chemical-signalling molecules, usually small proteins or steroids, secreted by endocrine cells that act to control or regulate specific physiological processes, including growth, development, metabolism, and reproduction. For example, insulin is a protein hormone that helps regulate the blood glucose level.
Table 4.1 Types and Functions of Proteins
| Type | Examples | Functions |
| Digestive enzymes | Amylase, lipase, pepsin, trypsin | Digest food by catabolizing nutrients into monomeric units |
| Transport | Hemoglobin, albumin | Carry substances in the blood or lymph throughout the body |
| Structural | Actin, tubulin, keratin | Mechanical support, like the cytoskeleton |
| Hormones | Insulin, thyroxine | Coordinate different body systems’ activities |
| Defence | Immunoglobulins | Protect the body from foreign pathogens |
| Contractile | Actin, myosin | Effect muscle contraction |
| Storage | Legume storage proteins, egg white (albumin) | Provide nourishment to the seedling and during embryonic development |
Amino Acids
Amino acids are the monomers that comprise proteins. Different arrangements of the same 20 types of amino acids comprise all proteins. Two rare new amino acids were discovered recently (selenocysteine and pyrrolysine), and additional new discoveries may be added to the list. Each amino acid has the same fundamental structure, which consists of a central carbon atom, or the alpha (α) carbon, bonded to an amino group (-NH2), a carboxyl group (-COOH), and to a hydrogen atom. Every amino acid also has another atom or group of atoms bonded to the central atom known as the R group (Figure 4.20).

Scientists use the name “amino acid” because these molecules contain both a basic amine group and an acidic carboxyl group in their general structure. As we mentioned, there are 20 common amino acids present in proteins. Nine of these are essential amino acids in humans because the human body cannot produce them and we obtain them from our diet. For each amino acid, the R group (or side chain) is different (Figure 4.21).
VISUAL CONNECTION
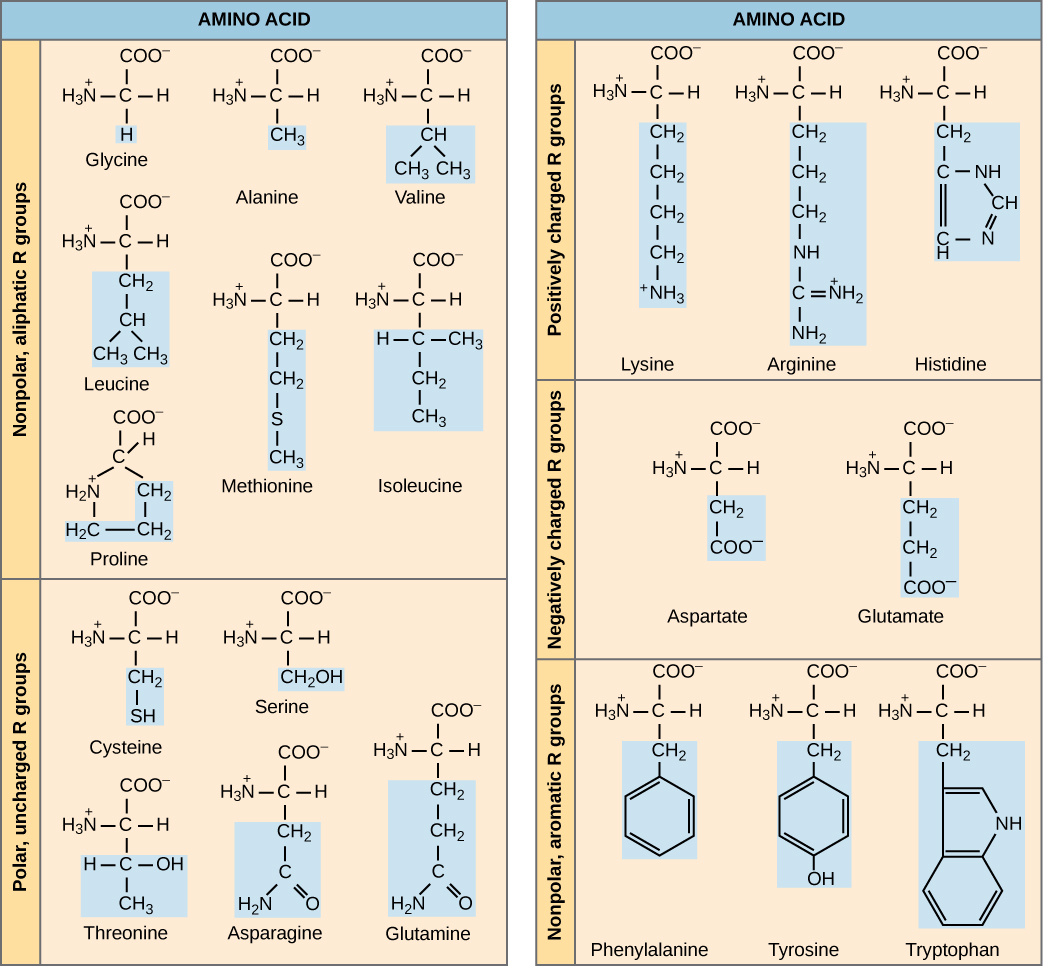
Which categories of amino acid would you expect to find on a soluble protein’s surface and which would you expect to find in the interior? What distribution of amino acids would you expect to find in a protein embedded in a lipid bilayer?
The chemical nature of the side chain determines the amino acid’s nature, that is, whether the amino acid is acidic, basic, polar, or nonpolar. For example, the amino acid glycine has a hydrogen atom as the R group. Amino acids such as valine, methionine, and alanine are nonpolar or hydrophobic in nature, while amino acids such as serine, threonine, and cysteine are polar and have hydrophilic side chains. Lysine and arginine are basic amino acids because their side chains can accept protons to become positively charged. Aspartic acid and glutamic acid have carboxyl groups in their side chains that deprotonate to give the negatively charged conjugate base aspartate and glutamate. Proline has an R group that is linked to the alpha amino group, forming a ring structure and is an exception to the amino acid’s standard structure.
A single uppercase letter or a three-letter abbreviation represents amino acids. For example, the letter V or the three-letter symbol Val represents valine.
Just as some fatty acids are essential to a diet, some amino acids also are necessary. These essential amino acids in humans include isoleucine, leucine, and cysteine. Essential amino acids refer to those necessary to build proteins in the body, but not those that the body produces. Which amino acids are essential varies from organism to organism.
The sequence and the number of amino acids ultimately determine the protein’s shape, size, and function. A covalent bond, or peptide bond, attaches to each amino acid, which a dehydration reaction forms. One amino acid’s carboxyl group and the incoming amino acid’s amino group combine, releasing a water molecule. The resulting bond is the peptide bond (Figure 4.22).
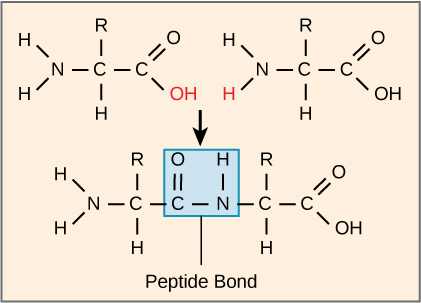
The products that such linkages form are peptides. As more amino acids join to this growing chain, the resulting chain is a polypeptide. Each polypeptide has a free amino group at one end. This end is the N terminal, or the amino terminal, and the other end has a free carboxyl group, also the C or carboxyl terminal. While the terms polypeptide and protein are sometimes used interchangeably, a polypeptide is technically a polymer of amino acids, whereas the term protein is used for a polypeptide or polypeptides that have combined together, often have bound non-peptide prosthetic groups, have a distinct shape, and have a unique function. After protein synthesis (translation), most proteins are modified. These are known as post-translational modifications. They may undergo cleavage, phosphorylation, or may require adding other chemical groups. Only after these modifications is the protein completely functional.
LINK TO LEARNING
Click through the steps of protein synthesis in this interactive tutorial.
EVOLUTION CONNECTION
The Evolutionary Significance of Cytochrome c
Cytochrome c is an important component of the electron transport chain, a part of cellular respiration, and it is normally located in the cellular organelle, the mitochondrion. This protein has a heme prosthetic group, and the heme’s central ion alternately reduces and oxidizes during electron transfer. Because this essential protein’s role in producing cellular energy is crucial, it has changed very little over millions of years. Protein sequencing has shown that there is a considerable amount of cytochrome c amino acid sequence homology among different species. In other words, we can assess evolutionary kinship by measuring the similarities or differences among various species’ DNA or protein sequences.
Scientists have determined that human cytochrome c contains 104 amino acids. For each cytochrome c molecule from different organisms that scientists have sequenced to date, 37 of these amino acids appear in the same position in all cytochrome c samples. This indicates that there may have been a common ancestor. On comparing the human and chimpanzee protein sequences, scientists did not find a sequence difference. When researchers compared human and rhesus monkey sequences, the single difference was in one amino acid. In another comparison, human to yeast sequencing shows a difference in the 44th position.
Protein Structure
As we discussed earlier, a protein’s shape is critical to its function. For example, an enzyme can bind to a specific substrate at an active site. If this active site is altered because of local changes or changes in overall protein structure, the enzyme may be unable to bind to the substrate. To understand how the protein gets its final shape or conformation, we need to understand the four levels of protein structure: primary, secondary, tertiary, and quaternary.
Primary Structure
Amino acids’ unique sequence in a polypeptide chain is its primary structure. For example, the pancreatic hormone insulin has two polypeptide chains, A and B, and they are linked together by disulfide bonds. The N terminal amino acid of the A chain is glycine; whereas, the C terminal amino acid is asparagine (Figure 4.23). The amino acid sequences in the A and B chains are unique to insulin.
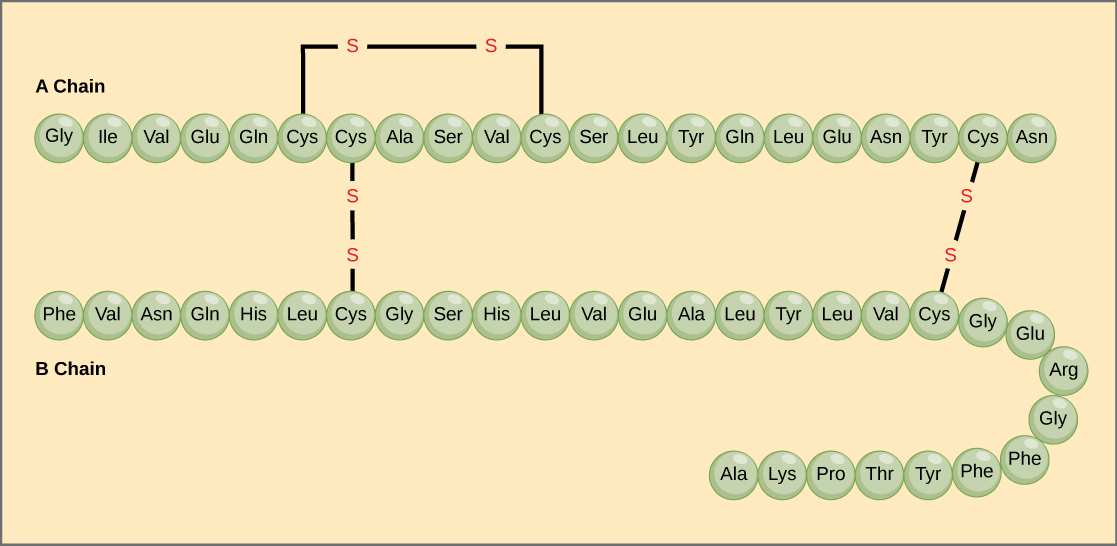
The gene encoding the protein ultimately determines the unique sequence for every protein. A change in nucleotide sequence of the gene’s coding region may lead to adding a different amino acid to the growing polypeptide chain, causing a change in protein structure and function. In sickle cell anemia, the hemoglobin β chain (a small portion of which we show in Figure 4.24) has a single amino acid substitution, causing a change in protein structure and function. Specifically, valine in the β chain substitutes the glutamic acid. What is most remarkable to consider is that a hemoglobin molecule is comprised of two alpha and two beta chains that each consist of about 150 amino acids. The molecule, therefore, has about 600 amino acids. The structural difference between a normal hemoglobin molecule and a sickle cell molecule—which dramatically decreases life expectancy—is a single amino acid of the 600. What is even more remarkable is that three nucleotides each encode those 600 amino acids, and a single base change (point mutation), 1 in 1800 bases causes the mutation.
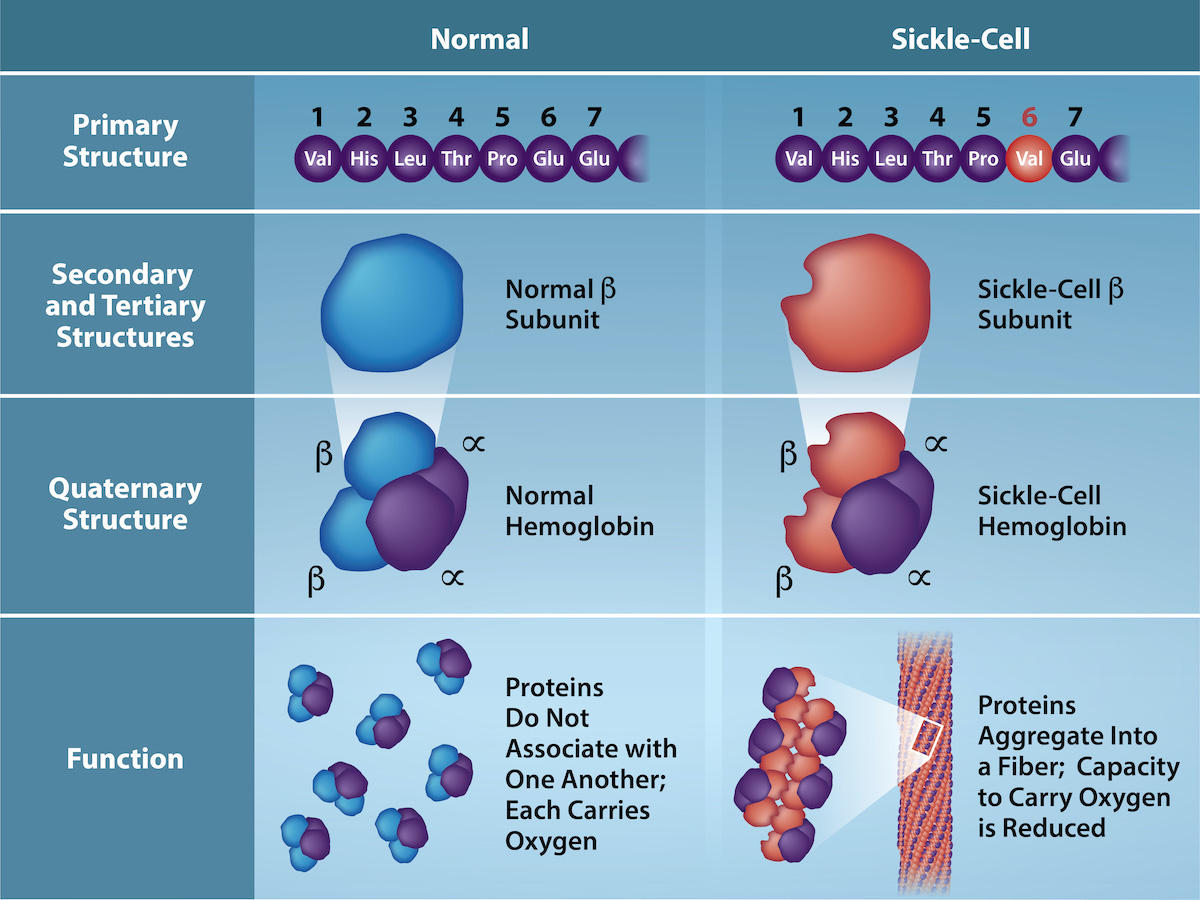
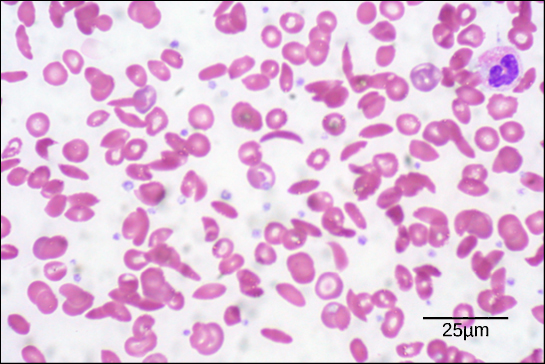
Because of this change of one amino acid in the chain, hemoglobin molecules form long fibers that distort the biconcave, or disc-shaped, red blood cells and causes them to assume a crescent or “sickle” shape, which clogs blood vessels (Figure 4.25). This can lead to myriad serious health problems such as breathlessness, dizziness, headaches, and abdominal pain for those affected by this disease. William Warrick Cardozo showed that sickle-cell anemia is an inherited disorder, meaning that the difference in the specific gene’s encoding region is passed down from parents to children. As you will learn in the genetics unit, the inheritance of such traits is determined by a combination of genes from both parents, and these very small differences can have significant impacts on organisms.
Secondary Structure
The local folding of the polypeptide in some regions gives rise to the secondary structure of the protein. The most common are the α-helix and β-pleated sheet structures (Figure 4.26). Both structures are held in shape by hydrogen bonds.
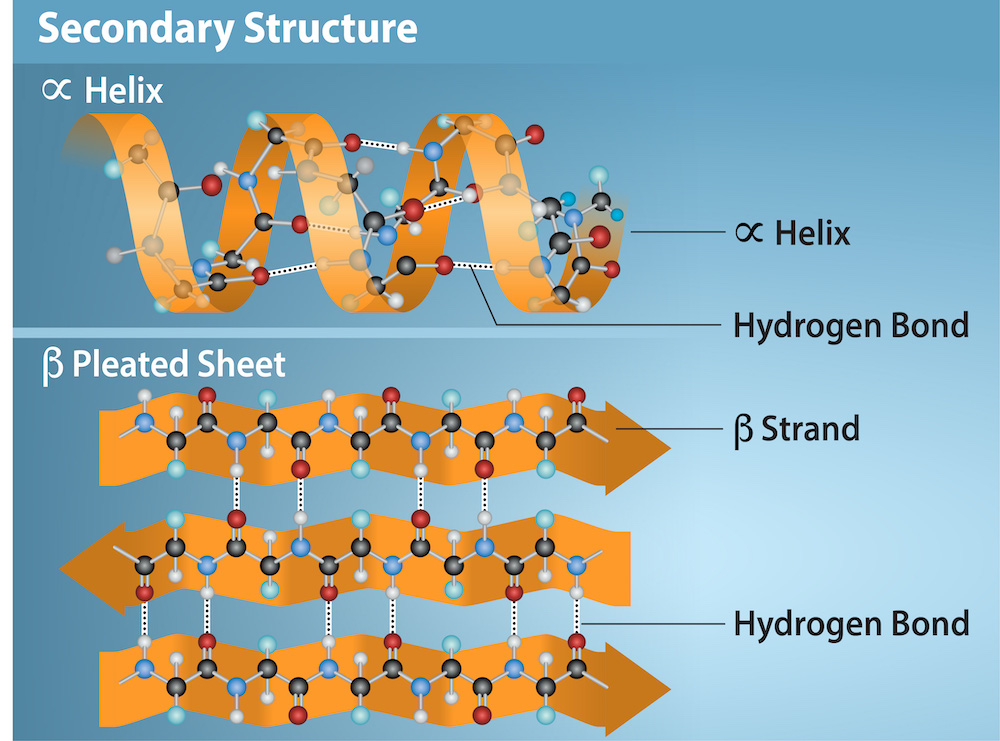
In the α-helix, the hydrogen bonds form between the oxygen atom in the carbonyl group in one amino acid and another amino acid that is four amino acids farther along the chain. Every helical turn in an alpha helix has 3.6 amino acid residues. The polypeptide’s R groups (the variant groups) protrude out from the α-helix chain.
In the β-pleated sheet, hydrogen bonding between atoms on the polypeptide chain’s backbone form the “pleats”. The R groups are attached to the carbons and extend above and below the pleat’s folds. The pleated segments align parallel or antiparallel to each other, and hydrogen bonds form between the partially positive hydrogen atom in the amino group and the partially negative oxygen atom in the peptide backbone’s carbonyl group. The α-helix and β-pleated sheet structures are in most globular and fibrous proteins and they play an important structural role.
Tertiary Structure
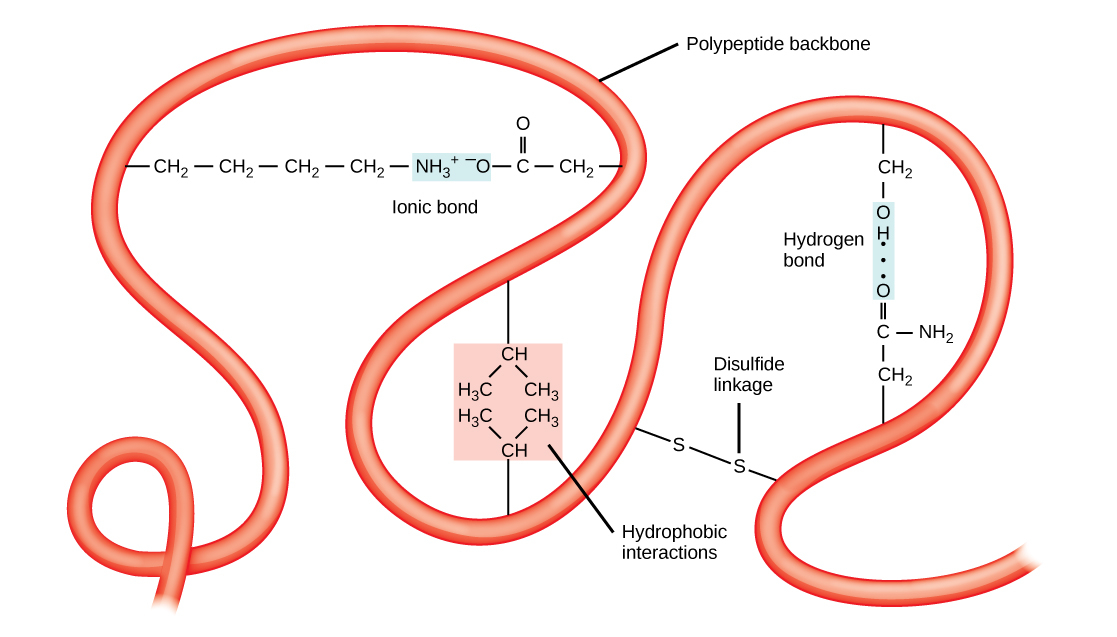
The polypeptide’s unique three-dimensional structure is its tertiary structure (Figure 4.27). This structure is in part due to chemical interactions at work on the polypeptide chain. Primarily, the interactions among R groups create the protein’s complex three-dimensional tertiary structure. The nature of the R groups in the amino acids involved can counteract the hydrogen bonds we described for standard secondary structures. For example, R groups with like charges repel each other and those with unlike charges are attracted to each other (ionic bonds). When protein folding takes place, the nonpolar amino acids’ hydrophobic R groups lie in the protein’s interior; whereas, the hydrophilic R groups lie on the outside. Scientists also call the former interaction types hydrophobic interactions. Interaction between cysteine side chains forms disulfide linkages in the presence of oxygen, the only covalent bond that forms during protein folding. All of these interactions, weak and strong, determine the protein’s final three-dimensional shape. When a protein loses its three-dimensional shape, it may no longer be functional.
Quaternary Structure
In nature, some proteins form from several polypeptides, or subunits, and the interaction of these subunits forms the quaternary structure. Weak interactions between the subunits help to stabilize the overall structure. For example, insulin (a globular protein) has a combination of hydrogen and disulfide bonds that cause it to mostly clump into a ball shape. Insulin starts out as a single polypeptide and loses some internal sequences in the presence of post-translational modification after forming the disulfide linkages that hold the remaining chains together. Silk (a fibrous protein), however, has a β-pleated sheet structure that is the result of hydrogen bonding between different chains. Figure 4.28 illustrates the four levels of protein structure (primary, secondary, tertiary, and quaternary).

Denaturation and Protein Folding
Each protein has its own unique sequence and shape that chemical interactions hold together. If the protein is subjected to changes in temperature, pH, or exposure to chemicals, the protein structure may change, losing its shape without losing its primary sequence in what scientists call denaturation. Denaturation is often reversible because the polypeptide’s primary structure is conserved in the process if the denaturing agent is removed, allowing the protein to resume its function. Sometimes denaturation is irreversible, leading to loss of function. One example of irreversible protein denaturation is frying an egg. The albumin protein in the liquid egg white denatures when placed in a hot pan. Not all proteins denature at high temperatures. For instance, Archaea that survive in hot springs have proteins that function at temperatures close to boiling. The stomach is also very acidic, has a low pH, and denatures proteins as part of the digestion process; however, the stomach’s digestive enzymes retain their activity under these conditions.
Protein folding is critical to its function. Scientists originally thought that the proteins themselves were responsible for the folding process. Only recently researchers discovered that often they receive assistance in the folding process from protein helpers, or chaperones (or chaperonins) that associate with the target protein during the folding process. They act by preventing polypeptide aggregation, and they disassociate from the protein once the target protein is folded.
Section Summary
Proteins are a class of macromolecules that perform a diverse range of functions for the cell. They help in metabolism by acting as enzymes, carriers, or hormones, and provide structural support. The building blocks of proteins (monomers) are amino acids. Each amino acid has a central carbon that bonds to an amino group, a carboxyl group, a hydrogen atom, and an R group or side chain. There are 20 commonly occurring amino acids, each of which differs in the R group. A peptide bond links each amino acid to its neighbors. A long amino acid chain is a polypeptide.
Proteins are organized at four levels: primary, secondary, tertiary, and (optional) quaternary. The primary structure is the amino acids’ unique sequence. The polypeptide’s local folding of the backbone forms structures such as the α-helix and β-pleated sheet constitutes the secondary structure. The overall three-dimensional structure is the tertiary structure. When two or more polypeptides combine to form the complete protein structure, the configuration is the protein’s quaternary structure. Protein shape and function are intricately linked. Any change in shape caused by changes in temperature or pH may lead to protein denaturation and a loss of function.
LINK TO LEARNING
For an additional perspective on proteins, view this animation called “Biomolecules: The Proteins.”
Exercises
Glossary
alpha (α)-helix: secondary protein structure formed by folding the polypeptide into a helix shape with hydrogen bonds stabilizing the structure
amino acid: a protein’s monomer; has a central carbon or alpha carbon to which an amino group, a carboxyl group, a hydrogen, and an R group or side chain is attached; the R group is different for all 20 common amino acids
beta (β)-pleated sheet: secondary protein structure in which hydrogen bonding forms “pleats” between atoms on the polypeptide chain’s backbone
chaperone (also, chaperonin): protein that helps nascent protein in the folding process
denaturation: loss of shape in a protein as a result of changes in temperature, pH, or chemical exposure
enzyme: catalyst in a biochemical reaction that is usually a complex or conjugated protein
hormone: chemical signaling molecule, usually protein or steroid, secreted by endocrine cells that act to control or regulate specific physiological processes
peptide bond: bond formed between two amino acids by a dehydration reaction
polypeptide: long chain of amino acids linked by peptide bonds
primary structure: linear sequence of amino acids in a protein
protein: biological macromolecule comprised of one or more amino acid chains
quaternary structure: association of discrete polypeptide subunits in a protein
secondary structure: regular structure that proteins form by intramolecular hydrogen bonding between the oxygen atom of one amino acid residue and the hydrogen attached to the nitrogen atom of another amino acid residue
tertiary structure: a protein’s three-dimensional conformation, including interactions between secondary structural elements; formed from interactions between amino acid side chains
Media Attributions
- Figure 4.24 by Rao, A., Tag, A. Ryan, K. and Fletcher, S. Department of Biology, Texas A&M University
- Figure 4.25 modification of work by Ed Uthman; scale-bar data from Matt Russell
- Figure 4.26 by Rao, A., Ryan, K. Fletcher, S. and Tag, A. Department of Biology, Texas A&M University
- Figure 4.28 by Rao, A. Ryan, K. and Tag, A. Department of Biology, Texas A&M University

