11.2 Dominant and Recessive Traits
Learning Objectives
By the end of this section, you will be able to:
- Explain the relationship between genotypes and phenotypes in dominant and recessive gene systems
- Use a Punnett square to calculate the expected proportions of genotypes and phenotypes in a monohybrid cross
- Explain Mendel’s law of segregation in terms of genetics and the events of meiosis
- Explain the purpose and methods of a test cross
The seven characteristics that Mendel evaluated in his pea plants were each expressed as one of two versions, or traits. Mendel deduced from his results that each individual had two discrete copies of the characteristic that are passed individually to offspring. We now call those two copies genes, which are carried on chromosomes. The reason we have two copies of each gene is that we inherit one from each parent. In fact, it is the chromosomes we inherit and the two copies of each gene are located on paired chromosomes. Recall that in meiosis these chromosomes are separated out into haploid gametes. This separation, or segregation, of the homologous chromosomes means also that only one of the copies of the gene gets moved into a gamete. The offspring are formed when that gamete unites with one from another parent and the two copies of each gene (and chromosome) are restored.
For cases in which a single gene controls a single characteristic, a diploid organism has two genetic copies that may or may not encode the same version of that characteristic. For example, one individual may carry a gene that determines white flower colour and a gene that determines violet flower colour. Gene variants that arise by mutation and exist at the same relative locations on homologous chromosomes are called alleles. Mendel examined the inheritance of genes with just two allele forms, but it is common to encounter more than two alleles for any given gene in a natural population.
Phenotypes and Genotypes
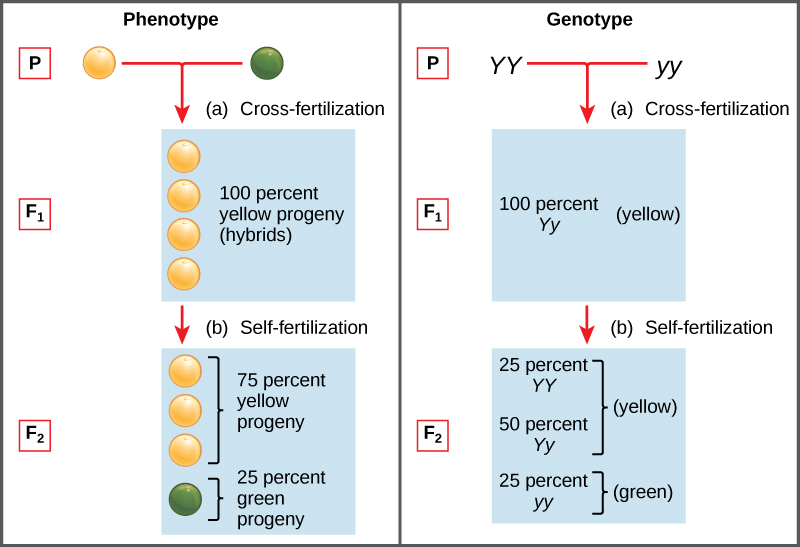
Two alleles for a given gene in a diploid organism are expressed and interact to produce physical characteristics. The observable traits expressed by an organism are referred to as its phenotype. An organism’s underlying genetic makeup, consisting of both the physically visible and the non-expressed alleles, is called its genotype. Mendel’s hybridization experiments demonstrate the difference between phenotype and genotype. For example, the phenotypes that Mendel observed in his crosses between pea plants with differing traits are connected to the diploid genotypes of the plants in the P, F1, and F2 generations. We will use a second trait that Mendel investigated, seed colour, as an example. Seed colour is governed by a single gene with two alleles. The yellow-seed allele is dominant and the green-seed allele is recessive. When true-breeding plants were cross-fertilized, in which one parent had yellow seeds and one had green seeds, all of the F1 hybrid offspring had yellow seeds. That is, the hybrid offspring were phenotypically identical to the true-breeding parent with yellow seeds. However, we know that the allele donated by the parent with green seeds was not simply lost because it reappeared in some of the F2 offspring (Figure 11.4). Therefore, the F1 plants must have been genotypically different from the parent with yellow seeds.
The pea plants that Mendel used in his experiments were each homozygous for the trait he was studying. Diploid organisms that are homozygous for a gene have two identical alleles, one on each of their homologous chromosomes. The genotype is often written as YY or yy, for which each letter represents one of the two alleles in the genotype. The dominant allele is capitalized and the recessive allele is lower case. The letter used for the gene (seed colour in this case) is usually related to the dominant trait (yellow allele, in this case, or “Y”). Mendel’s parental pea plants always bred true because both produced gametes carried the same allele. When pea plants with contrasting traits were cross-fertilized, all of the offspring were heterozygous for the contrasting trait, meaning their genotype had different alleles for the gene being examined. For example, the F1 yellow plants that received a Y allele from their yellow parent and a y allele from their green parent had the genotype Yy.
Law of Dominance
Our discussion of homozygous and heterozygous organisms brings us to why the F1 heterozygous offspring were identical to one of the parents, rather than expressing both alleles. In all seven pea-plant characteristics, one of the two contrasting alleles was dominant, and the other was recessive. Mendel called the dominant allele the expressed unit factor; the recessive allele was referred to as the latent unit factor. We now know that these so-called unit factors are actually genes on homologous chromosomes. For a gene that is expressed in a dominant and recessive pattern, homozygous dominant and heterozygous organisms will look identical (that is, they will have different genotypes but the same phenotype), and the recessive allele will only be observed in homozygous recessive individuals (Table 11.4).
| Homozygous | Heterozygous | Homozygous | |
|---|---|---|---|
| Genotype | YY | Yy | yy |
| Phenotype | yellow | yellow | green |
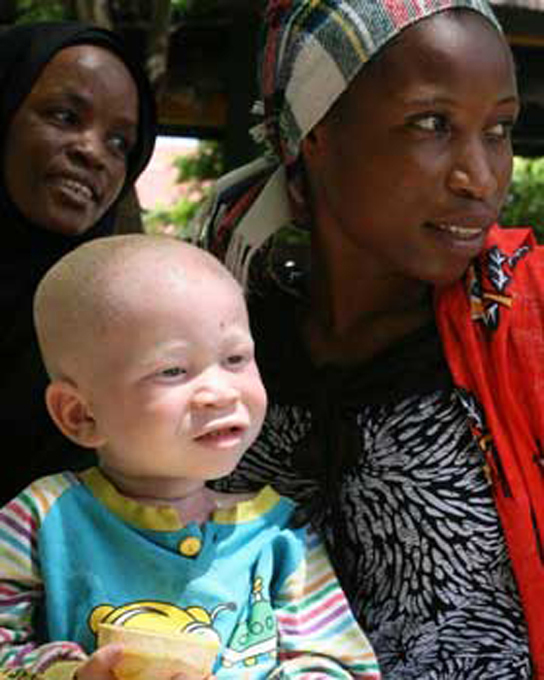
Mendel’s law of dominance states that in a heterozygote, one trait will conceal the presence of another trait for the same characteristic. For example, when crossing true-breeding violet-flowered plants with true-breeding white-flowered plants, all of the offspring were violet-flowered, even though they all had one allele for violet and one allele for white. Rather than both alleles contributing to a phenotype, the dominant allele will be expressed exclusively. The recessive allele will remain latent, but will be transmitted to offspring in the same manner as that by which the dominant allele is transmitted. The recessive trait will only be expressed by offspring that have two copies of this allele (Figure 11.5), and these offspring will breed true when self-crossed. The child in the photo expresses albinism, a recessive trait.

Equal Segregation of Alleles
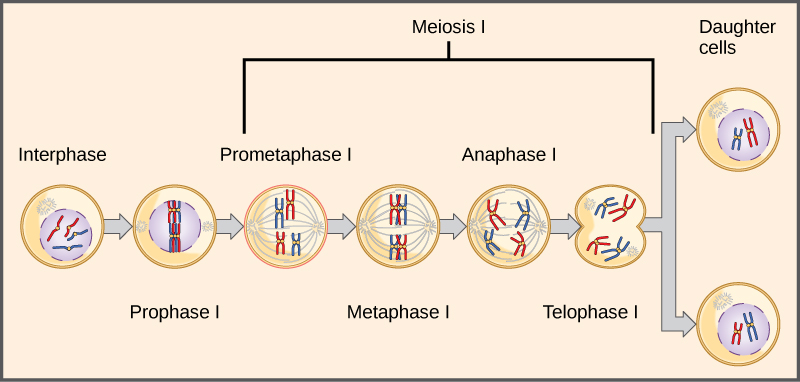
Observing that true-breeding pea plants with contrasting traits gave rise to F1 generations that all expressed the dominant trait and F2 generations that expressed the dominant and recessive traits in a 3:1 ratio, Mendel proposed the law of segregation. This law states that paired unit factors (genes) must segregate equally into gametes such that offspring have an equal likelihood of inheriting either factor. For the F2 generation of a monohybrid cross, the following three possible combinations of genotypes result: homozygous dominant, heterozygous, or homozygous recessive. Because heterozygotes could arise from two different pathways (receiving one dominant and one recessive allele from either parent), and because heterozygotes and homozygous dominant individuals are phenotypically identical, the law supports Mendel’s observed 3:1 phenotypic ratio. The equal segregation of alleles is the reason we can apply the Punnett square to accurately predict the offspring of parents with known genotypes. The physical basis of Mendel’s law of segregation is the first division of meiosis in which the homologous chromosomes with their different versions of each gene are segregated into daughter nuclei. The role of the meiotic segregation of chromosomes in sexual reproduction was not understood by the scientific community during Mendel’s lifetime (Figure 11.6).
Monohybrid Cross and the Punnett Square
When fertilization occurs between two true-breeding parents that differ by only the characteristic being studied, the process is called a monohybrid cross, and the resulting offspring are called monohybrids. Mendel performed seven types of monohybrid crosses, each involving contrasting traits for different characteristics. Out of these crosses, all of the F1 offspring had the phenotype of one parent, and the F2 offspring had a 3:1 phenotypic ratio. On the basis of these results, Mendel postulated that each parent in the monohybrid cross contributed one of two paired unit factors to each offspring, and every possible combination of unit factors was equally likely.
The results of Mendel’s research can be explained in terms of probabilities, which are mathematical measures of likelihood. The probability of an event is calculated by the number of times the event occurs divided by the total number of opportunities for the event to occur. A probability of one (100 percent) for some event indicates that it is guaranteed to occur, whereas a probability of zero (0 percent) indicates that it is guaranteed to not occur, and a probability of 0.5 (50 percent) means it has an equal chance of occurring or not occurring.
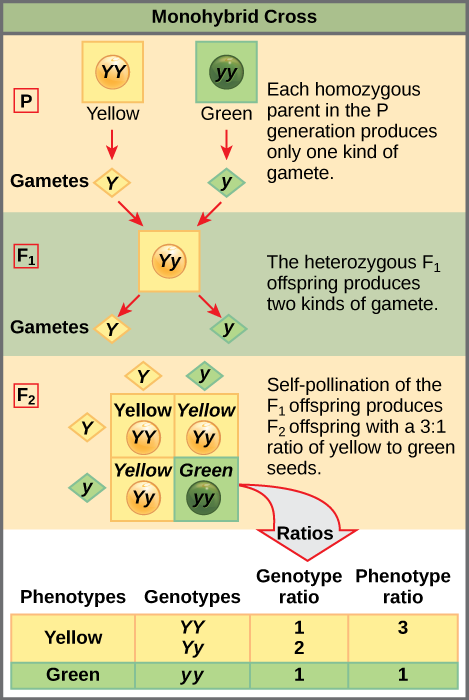
To demonstrate this with a monohybrid cross, consider the case of true-breeding pea plants with yellow versus green seeds. The dominant seed colour is yellow; therefore, the parental genotypes were YY for the plants with yellow seeds and yy for the plants with green seeds. A Punnett square, devised by the British geneticist Reginald Punnett, is useful for determining probabilities because it is drawn to predict all possible outcomes of all possible random fertilization events and their expected frequencies. Figure 11.7 shows a Punnett square for a cross between a plant with yellow peas and one with green peas. To prepare a Punnett square, all possible combinations of the parental alleles (the genotypes of the gametes) are listed along the top (for one parent) and side (for the other parent) of a grid. The combinations of egg and sperm gametes are then made in the boxes in the table on the basis of which alleles are combining. Each box then represents the diploid genotype of a zygote, or fertilized egg. Because each possibility is equally likely, genotypic ratios can be determined from a Punnett square. If the pattern of inheritance (dominant and recessive) is known, the phenotypic ratios can be inferred as well. For a monohybrid cross of two true-breeding parents, each parent contributes one type of allele. In this case, only one genotype is possible in the F1 offspring. All offspring are Yy and have yellow seeds.
When the F1 offspring are crossed with each other, each has an equal probability of contributing either a Y or a y to the F2 offspring. The result is a 1 in 4 (25 percent) probability of both parents contributing a Y, resulting in an offspring with a yellow phenotype; a 25 percent probability of parent A contributing a Y and parent B a y, resulting in offspring with a yellow phenotype; a 25 percent probability of parent A contributing a y and parent B a Y, also resulting in a yellow phenotype; and a (25 percent) probability of both parents contributing a y, resulting in a green phenotype. When counting all four possible outcomes, there is a 3 in 4 probability of offspring having the yellow phenotype and a 1 in 4 probability of offspring having the green phenotype. This explains why the results of Mendel’s F2 generation occurred in a 3:1 phenotypic ratio. Using large numbers of crosses, Mendel was able to calculate probabilities, found that they fit the model of inheritance, and use these to predict the outcomes of other crosses.
Test Cross
Beyond predicting the offspring of a cross between known homozygous or heterozygous parents, Mendel also developed a way to determine whether an organism that expressed a dominant trait was a heterozygote or a homozygote. Called the test cross, this technique is still used by plant and animal breeders. In a test cross, the dominant-expressing organism is crossed with an organism that is homozygous recessive for the same characteristic. If the dominant-expressing organism is a homozygote, then all F1 offspring will be heterozygotes expressing the dominant trait (Figure 11.7). Alternatively, if the dominant-expressing organism is a heterozygote, the F1 offspring will exhibit a 1:1 ratio of heterozygotes and recessive homozygotes (Figure 11.8). The test cross further validates Mendel’s postulate that pairs of unit factors segregate equally.
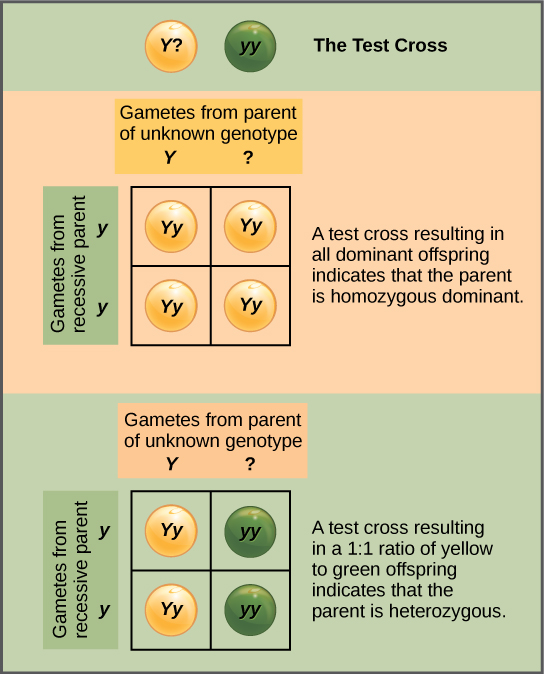
In pea plants, round peas (R) are dominant to wrinkled peas (r). You do a test cross between a pea plant with wrinkled peas (genotype rr) and a plant of unknown genotype that has round peas. You end up with three plants, all which have round peas. From this data, can you tell if the parent plant is homozygous dominant or heterozygous?
You cannot be sure if the plant is homozygous or heterozygous as the data set is too small: by random chance, all three plants might have acquired only the dominant gene even if the recessive one is present.
VISUAL CONNECTION
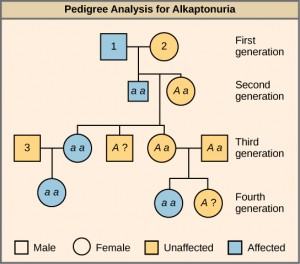
In this pedigree (Figure 11.9), individuals with the disorder are indicated in blue and have the genotype aa. Unaffected individuals are indicated in yellow and have the genotype AA or Aa. Note that it is often possible to determine a person’s genotype from the genotype of their offspring. For example, if neither parent has the disorder but their child does, they must be heterozygous. Two individuals on the pedigree have an unaffected phenotype but unknown genotype. Because they do not have the disorder, they must have at least one normal allele, so their genotype gets the “A?” designation.
What are the genotypes of the individuals labeled 1, 2 and 3?
Alternatives to Dominance and Recessiveness
Mendel’s experiments with pea plants suggested that: (1) two “units” or alleles exist for every gene; (2) alleles maintain their integrity in each generation (no blending); and (3) in the presence of the dominant allele, the recessive allele is hidden and makes no contribution to the phenotype. Therefore, recessive alleles can be “carried” and not expressed by individuals. Such heterozygous individuals are sometimes referred to as “carriers.” Further genetic studies in other plants and animals have shown that much more complexity exists, but that the fundamental principles of Mendelian genetics still hold true. In the sections to follow, we consider some of the extensions of Mendelism. If Mendel had chosen an experimental system that exhibited these genetic complexities, it’s possible that he would not have understood what his results meant.
Incomplete Dominance
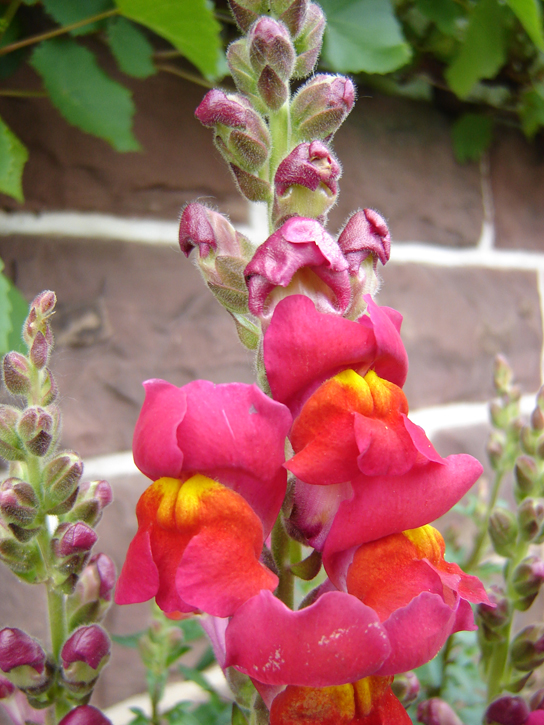
Mendel’s results, that traits are inherited as dominant and recessive pairs, contradicted the view at that time that offspring exhibited a blend of their parents’ traits. However, the heterozygote phenotype occasionally does appear to be intermediate between the two parents. For example, in the snapdragon, Antirrhinum majus (Figure 11.10), a cross between a homozygous parent with white flowers (CWCW) and a homozygous parent with red flowers (CRCR) will produce offspring with pink flowers (CRCW). (Note that different genotypic abbreviations are used for Mendelian extensions to distinguish these patterns from simple dominance and recessiveness.) This pattern of inheritance is described as incomplete dominance, denoting the expression of two contrasting alleles such that the individual displays an intermediate phenotype. The allele for red flowers is incompletely dominant over the allele for white flowers. However, the results of a heterozygote self-cross can still be predicted, just as with Mendelian dominant and recessive crosses. In this case, the genotypic ratio would be 1 CRCR:2 CRCW:1 CWCW, and the phenotypic ratio would be 1:2:1 for red:pink:white.

Codominance
A variation on incomplete dominance is codominance, in which both alleles for the same characteristic are simultaneously expressed in the heterozygote. An example of codominance is the MN blood groups of humans. The M and N alleles are expressed in the form of an M or N antigen present on the surface of red blood cells. Homozygotes (LMLM and LNLN) express either the M or the N allele, and heterozygotes (LMLN) express both alleles equally. In a self-cross between heterozygotes expressing a codominant trait, the three possible offspring genotypes are phenotypically distinct. However, the 1:2:1 genotypic ratio characteristic of a Mendelian monohybrid cross still applies.
Multiple Alleles
Mendel implied that only two alleles, one dominant and one recessive, could exist for a given gene. We now know that this is an oversimplification. Although individual humans (and all diploid organisms) can only have two alleles for a given gene, multiple alleles may exist at the population level such that many combinations of two alleles are observed. Note that when many alleles exist for the same gene, the convention is to denote the most common phenotype or genotype among wild animals as the wild type (often abbreviated “+”); this is considered the standard or norm. All other phenotypes or genotypes are considered variants of this standard, meaning that they deviate from the wild type. The variant may be recessive or dominant to the wild-type allele.
An example of multiple alleles is coat colour in rabbits (Figure 11.11). Here, four alleles exist for the c gene. The wild-type version, C+C+, is expressed as brown fur. The chinchilla phenotype, cchcch, is expressed as black-tipped white fur. The Himalayan phenotype, chch, has black fur on the extremities and white fur elsewhere. Finally, the albino, or “colourless” phenotype, cc, is expressed as white fur. In cases of multiple alleles, dominance hierarchies can exist. In this case, the wild-type allele is dominant over all the others, chinchilla is incompletely dominant over Himalayan and albino, and Himalayan is dominant over albino. This hierarchy, or allelic series, was revealed by observing the phenotypes of each possible heterozygote offspring.
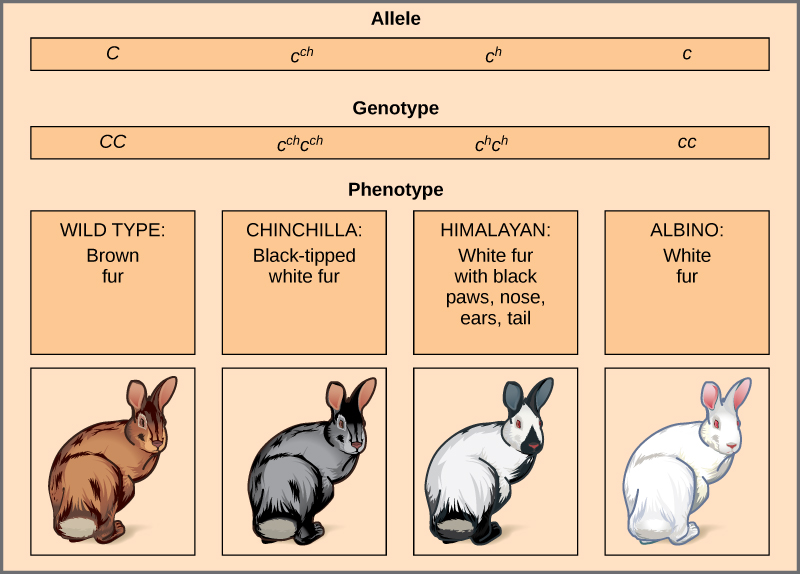
The complete dominance of a wild-type phenotype over all other mutants often occurs as an effect of “dosage” of a specific gene product, such that the wild-type allele supplies the correct amount of gene product whereas the mutant alleles cannot. For the allelic series in rabbits, the wild-type allele may supply a given dosage of fur pigment, whereas the mutants supply a lesser dosage or none at all. Interestingly, the Himalayan phenotype is the result of an allele that produces a temperature-sensitive gene product that only produces pigment in the cooler extremities of the rabbit’s body.
Alternatively, one mutant allele can be dominant over all other phenotypes, including the wild type. This may occur when the mutant allele somehow interferes with the genetic message so that even a heterozygote with one wild-type allele copy expresses the mutant phenotype. One way in which the mutant allele can interfere is by enhancing the function of the wild-type gene product or changing its distribution in the body. One example of this is the Antennapedia mutation in Drosophila (Figure 11.12). In this case, the mutant allele expands the distribution of the gene product, and as a result, the Antennapedia heterozygote develops legs on its head where its antennae should be.

Multiple Alleles Confer Drug Resistance in the Malaria Parasite
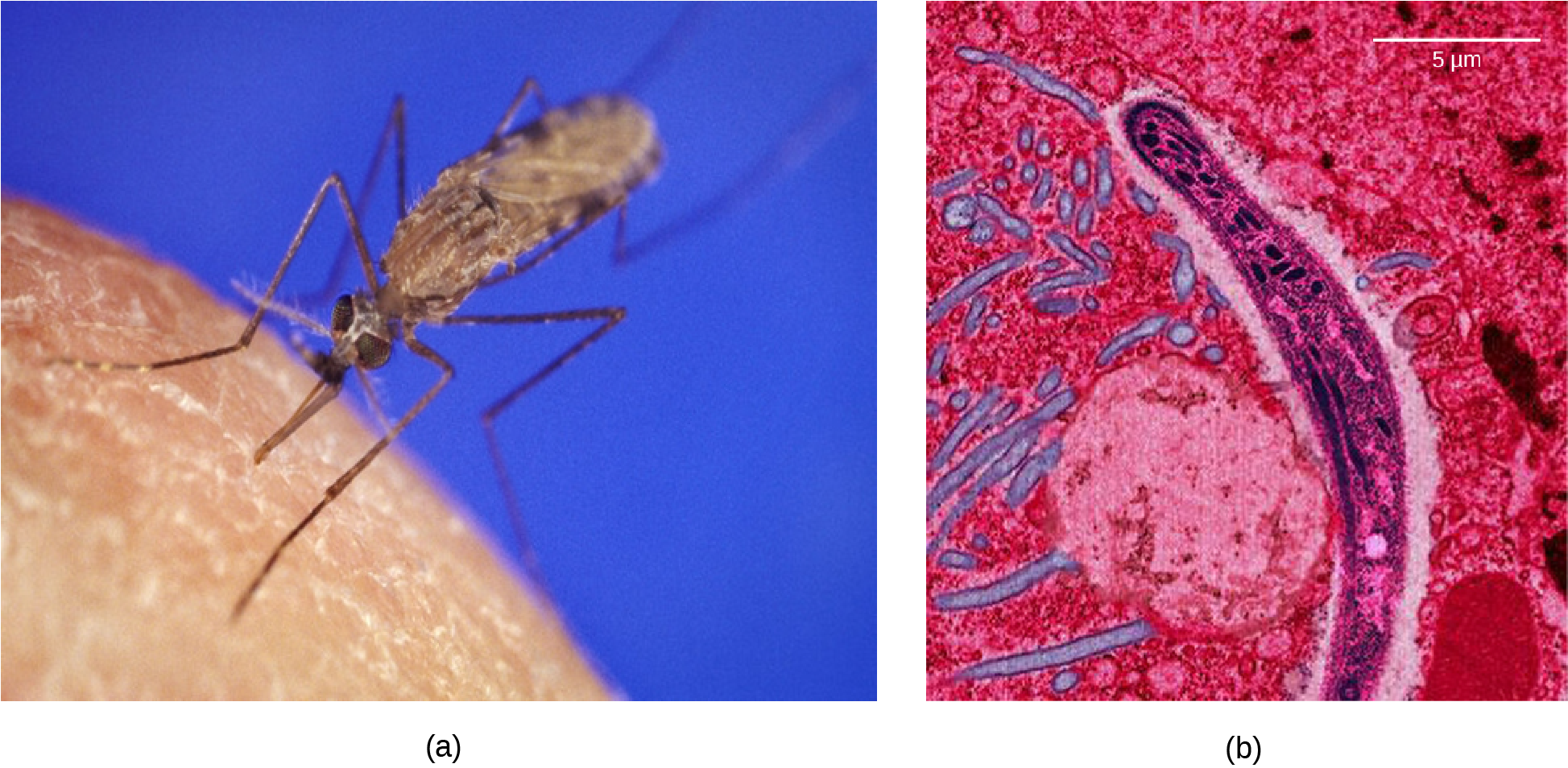
Malaria is a parasitic disease in humans that is transmitted by infected female mosquitoes, including Anopheles gambiae (Figure 11.13a), and is characterized by cyclic high fevers, chills, flu-like symptoms, and severe anemia. Plasmodium falciparum and P. vivax are the most common causative agents of malaria, and P. falciparum is the most deadly (Figure 11.13b). When promptly and correctly treated, P. falciparum malaria has a mortality rate of 0.1 percent. However, in some parts of the world, the parasite has evolved resistance to commonly used malaria treatments, so the most effective malarial treatments can vary by geographic region.
In Southeast Asia, Africa, and South America, P. falciparum has developed resistance to the anti-malarial drugs chloroquine, mefloquine, and sulfadoxine-pyrimethamine. P. falciparum, which is haploid during the life stage in which it is infectious to humans, has evolved multiple drug-resistant mutant alleles of the dhps gene. Varying degrees of sulfadoxine resistance are associated with each of these alleles. Being haploid, P. falciparum needs only one drug-resistant allele to express this trait.
In Southeast Asia, different sulfadoxine-resistant alleles of the dhps gene are localized to different geographic regions. This is a common evolutionary phenomenon that occurs because drug-resistant mutants arise in a population and interbreed with other P. falciparum isolates in close proximity. Sulfadoxine-resistant parasites cause considerable human hardship in regions where this drug is widely used as an over-the-counter malaria remedy. As is common with pathogens that multiply to large numbers within an infection cycle, P. falciparum evolves relatively rapidly (over a decade or so) in response to the selective pressure of commonly used anti-malarial drugs. For this reason, scientists must constantly work to develop new drugs or drug combinations to combat the worldwide malaria burden.1
X-Linked Traits
In humans, as well as in many other animals and some plants, the sex of the individual is determined by sex chromosomes. The sex chromosomes are one pair of non-homologous chromosomes. Until now, we have only considered inheritance patterns among non-sex chromosomes, or autosomes. In addition to 22 homologous pairs of autosomes, human females have a homologous pair of X chromosomes, whereas human males have an XY chromosome pair. Although the Y chromosome contains a small region of similarity to the X chromosome so that they can pair during meiosis, the Y chromosome is much shorter and contains many fewer genes. When a gene being examined is present on the X chromosome, but not on the Y chromosome, it is said to be X-linked.
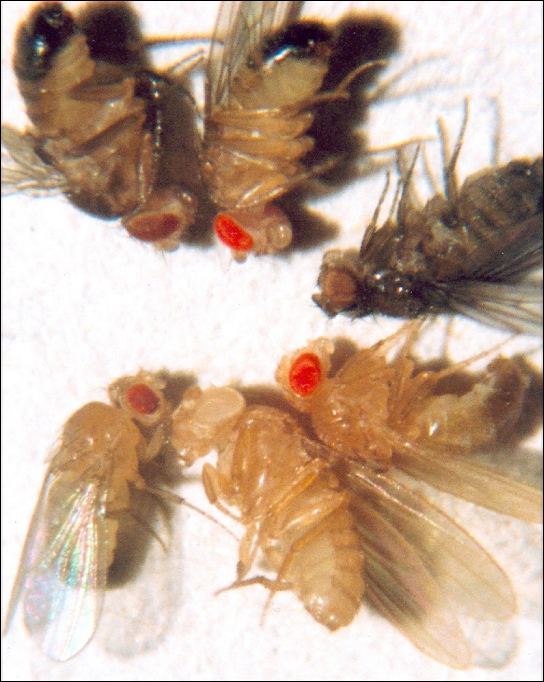
Eye colour in Drosophila was one of the first X-linked traits to be identified. Thomas Hunt Morgan mapped this trait to the X chromosome in 1910. Like humans, Drosophila males have an XY chromosome pair, and females are XX. In flies, the wild-type eye colour is red (XW) and it is dominant to white eye colour (Xw) (Figure 11.14). Because of the location of the eye-colour gene, reciprocal crosses do not produce the same offspring ratios. Males are said to be hemizygous, because they have only one allele for any X-linked characteristic. Hemizygosity makes the descriptions of dominance and recessiveness irrelevant for XY males. Drosophila males lack a second allele copy on the Y chromosome; that is, their genotype can only be XWY or XwY. In contrast, females have two allele copies of this gene and can be XWXW, XWXw, or XwXw.

In an X-linked cross, the genotypes of F1 and F2 offspring depend on whether the recessive trait was expressed by the male or the female in the P1 generation. With regard to Drosophila eye colour, when the P1 male expresses the white-eye phenotype and the female is homozygous red-eyed, all members of the F1 generation exhibit red eyes (Figure 11.15). The F1 females are heterozygous (XWXw), and the males are all XWY, having received their X chromosome from the homozygous dominant P1 female and their Y chromosome from the P1 male. A subsequent cross between the XWXw female and the XWY male would produce only red-eyed females (with XWXW or XWXw genotypes) and both red- and white-eyed males (with XWY or XwY genotypes). Now, consider a cross between a homozygous white-eyed female and a male with red eyes. The F1 generation would exhibit only heterozygous red-eyed females (XWXw) and only white-eyed males (XwY). Half of the F2 females would be red-eyed (XWXw) and half would be white-eyed (XwXw). Similarly, half of the F2 males would be red-eyed (XWY) and half would be white-eyed (XwY).
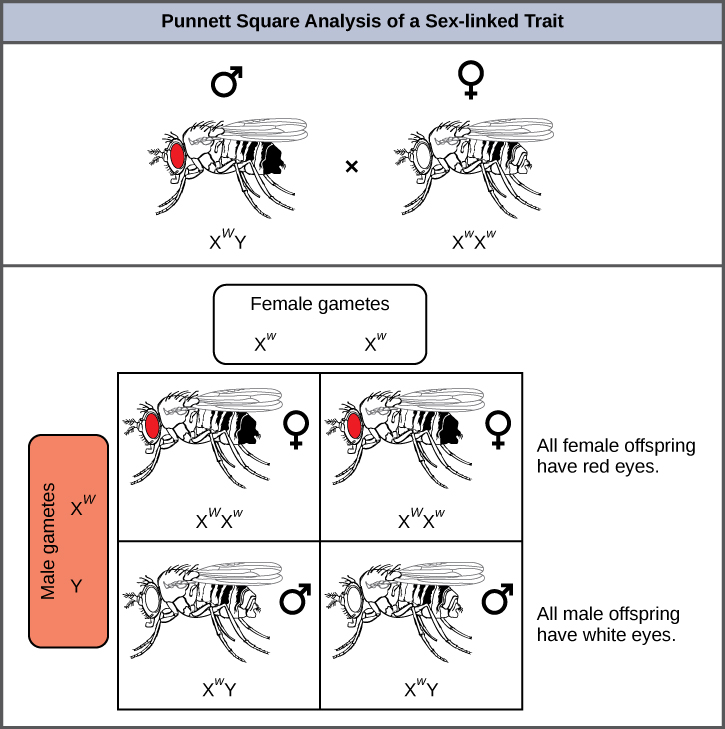
What ratio of offspring would result from a cross between a white-eyed male and a female that is heterozygous for red eye colour?
Discoveries in fruit fly genetics can be applied to human genetics. When a female parent is homozygous for a recessive X-linked trait, she will pass the trait on to 100 percent of her offspring. Her male offspring are, therefore, destined to express the trait, as they will inherit their father’s Y chromosome. In humans, the alleles for certain conditions (some forms of colour blindness, hemophilia, and muscular dystrophy) are X-linked. Females who are heterozygous for these diseases are said to be carriers and may not exhibit any phenotypic effects. These females will pass the disease to half of their sons and will pass carrier status to half of their daughters; therefore, recessive X-linked traits appear more frequently in males than females.
In some groups of organisms with sex chromosomes, the sex with the non-homologous sex chromosomes is the female rather than the male. This is the case for all birds. In this case, sex-linked traits will be more likely to appear in the female, in which they are hemizygous.
Human Sex-linked Disorders
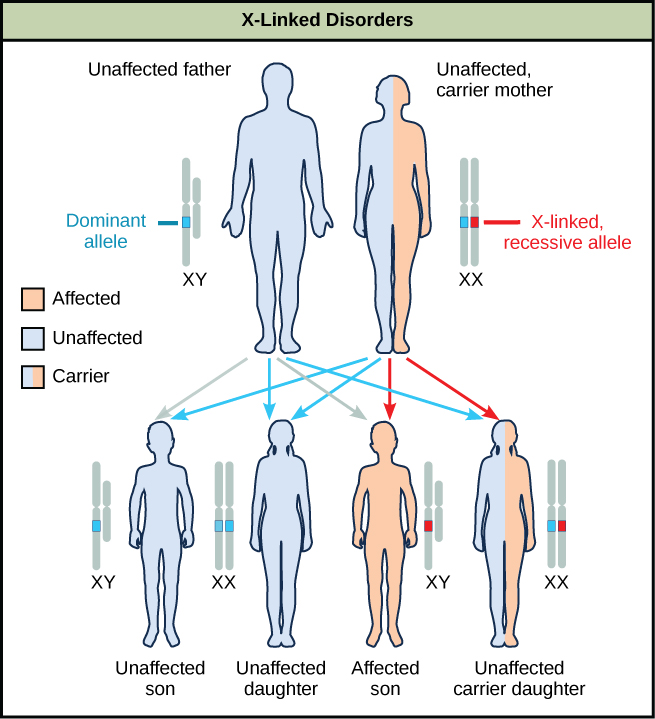
Sex-linkage studies in Morgan’s laboratory provided the fundamentals for understanding X-linked recessive disorders in humans, which include red-green colour blindness, and Types A and B hemophilia. Because human males need to inherit only one recessive mutant X allele to be affected, X-linked disorders are disproportionately observed in males. Females must inherit recessive X-linked alleles from both of their parents in order to express the trait. When they inherit one recessive X-linked mutant allele and one dominant X-linked wild-type allele, they are carriers of the trait and are typically unaffected. Carrier females can manifest mild forms of the trait due to the inactivation of the dominant allele located on one of the X chromosomes. However, female carriers can contribute the trait to their sons, resulting in the son exhibiting the trait, or they can contribute the recessive allele to their daughters, resulting in the daughters being carriers of the trait (Figure 11.16). Although some Y-linked recessive disorders exist, typically they are associated with infertility in males and are therefore not transmitted to subsequent generations.
Watch this video to learn more about sex-linked traits.
Lethality
A large proportion of genes in an individual’s genome are essential for survival. Occasionally, a nonfunctional allele for an essential gene can arise by mutation and be transmitted in a population as long as individuals with this allele also have a wild-type, functional copy. The wild-type allele functions at a capacity sufficient to sustain life and is therefore considered to be dominant over the nonfunctional allele. However, consider two heterozygous parents that have a genotype of wild-type/nonfunctional mutant for a hypothetical essential gene. In one quarter of their offspring, we would expect to observe individuals that are homozygous recessive for the nonfunctional allele. Because the gene is essential, these individuals might fail to develop past fertilization, die in utero, or die later in life, depending on what life stage requires this gene. An inheritance pattern in which an allele is only lethal in the homozygous form and in which the heterozygote may be normal or have some altered nonlethal phenotype is referred to as recessive lethal.
For crosses between heterozygous individuals with a recessive lethal allele that causes death before birth when homozygous, only wild-type homozygotes and heterozygotes would be observed. The genotypic ratio would therefore be 2:1. In other instances, the recessive lethal allele might also exhibit a dominant (but not lethal) phenotype in the heterozygote. For instance, the recessive lethal Curly allele in Drosophila affects wing shape in the heterozygote form but is lethal in the homozygote.
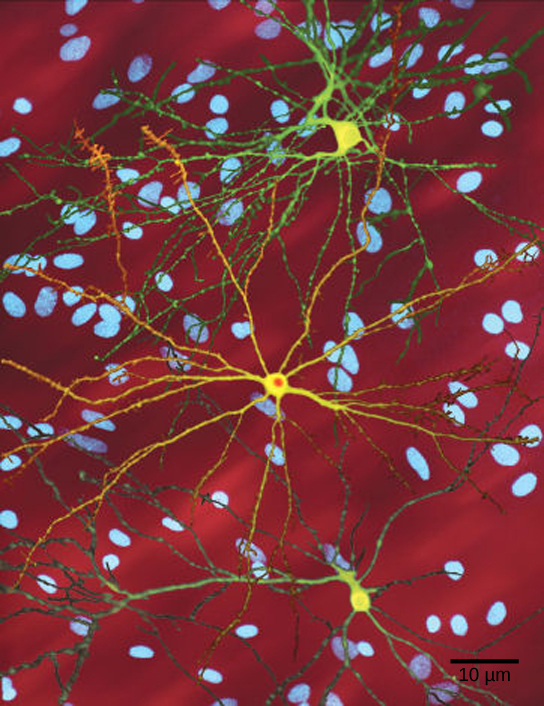
A single copy of the wild-type allele is not always sufficient for normal functioning or even survival. The dominant lethal inheritance pattern is one in which an allele is lethal both in the homozygote and the heterozygote; this allele can only be transmitted if the lethality phenotype occurs after reproductive age. Individuals with mutations that result in dominant lethal alleles fail to survive even in the heterozygote form. Dominant lethal alleles are very rare because, as you might expect, the allele only lasts one generation and is not transmitted. However, just as the recessive lethal allele might not immediately manifest the phenotype of death, dominant lethal alleles also might not be expressed until adulthood. Once the individual reaches reproductive age, the allele may be unknowingly passed on, resulting in a delayed death in both generations. An example of this in humans is Huntington’s disease, in which the nervous system gradually wastes away (Figure 11.17). People who are heterozygous for the dominant Huntington allele (Hh) will inevitably develop the fatal disease. However, the onset of Huntington’s disease may not occur until age 40, at which point the afflicted persons may have already passed the allele to 50 percent of their offspring.
Section Summary
When true-breeding, or homozygous, individuals that differ for a certain trait are crossed, all of the offspring will be heterozygous for that trait. If the traits are inherited as dominant and recessive, the F1 offspring will all exhibit the same phenotype as the parent homozygous for the dominant trait. If these heterozygous offspring are self-crossed, the resulting F2 offspring will be equally likely to inherit gametes carrying the dominant or recessive trait, giving rise to offspring of which one quarter are homozygous dominant, half are heterozygous, and one quarter are homozygous recessive. Because homozygous dominant and heterozygous individuals are phenotypically identical, the observed traits in the F2 offspring will exhibit a ratio of three dominant to one recessive.
Mendel postulated that genes (characteristics) are inherited as pairs of alleles (traits) that behave in a dominant and recessive pattern. Alleles segregate into gametes such that each gamete is equally likely to receive either one of the two alleles present in a diploid individual. In addition, genes are assorted into gametes independently of one another. That is, in general, alleles are not more likely to segregate into a gamete with a particular allele of another gene.
Alleles do not always behave in dominant and recessive patterns. Incomplete dominance describes situations in which the heterozygote exhibits a phenotype that is intermediate between the homozygous phenotypes. Codominance describes the simultaneous expression of both of the alleles in the heterozygote. Although diploid organisms can only have two alleles for any given gene, it is common for more than two alleles of a gene to exist in a population. In humans, as in many animals and some plants, females have two X chromosomes and males have one X and one Y chromosome. Genes that are present on the X but not the Y chromosome are said to be X-linked, such that males only inherit one allele for the gene, and females inherit two. Finally, some alleles can be lethal. Recessive lethal alleles are only lethal in homozygotes, but dominant lethal alleles are fatal in heterozygotes as well.
Exercises
Glossary
allele: one of two or more variants of a gene that determines a particular trait for a characteristic
dihybrid: the result of a cross between two true-breeding parents that express different traits for two characteristics
genotype: the underlying genetic makeup, consisting of both physically visible and non-expressed alleles, of an organism
heterozygous: having two different alleles for a given gene on the homologous chromosomes
homozygous: having two identical alleles for a given gene on the homologous chromosomes
law of dominance: in a heterozygote, one trait will conceal the presence of another trait for the same characteristic
law of independent assortment: genes do not influence each other with regard to sorting of alleles into gametes; every possible combination of alleles is equally likely to occur
law of segregation: paired unit factors (i.e., genes) segregate equally into gametes such that offspring have an equal likelihood of inheriting any combination of factors
monohybrid: the result of a cross between two true-breeding parents that express different traits for only one characteristic
phenotype: the observable traits expressed by an organism
Punnett square: a visual representation of a cross between two individuals in which the gametes of each individual are denoted along the top and side of a grid, respectively, and the possible zygotic genotypes are recombined at each box in the grid
test cross: a cross between a dominant expressing individual with an unknown genotype and a homozygous recessive individual; the offspring phenotypes indicate whether the unknown parent is heterozygous or homozygous for the dominant trait
Media Attributions
- Figure 11.10 by storebukkebruse/Flickr
- Figure 11.13 by (a) James D. Gathany; (b) Ute Frevert; false colour by Margaret Shear; scale-bar data from Matt Russell
- Figure 11.17 by Dr. Steven Finkbeiner, Gladstone Institute of Neurological Disease, The Taube-Koret Center for Huntington’s Disease Research, and the University of California San Francisco/Wikimedia

